| # tokenizing DNA sequences | |
| Tokenizers for DNA tokenization for enigma-1.5b model. | |
| ## Overview | |
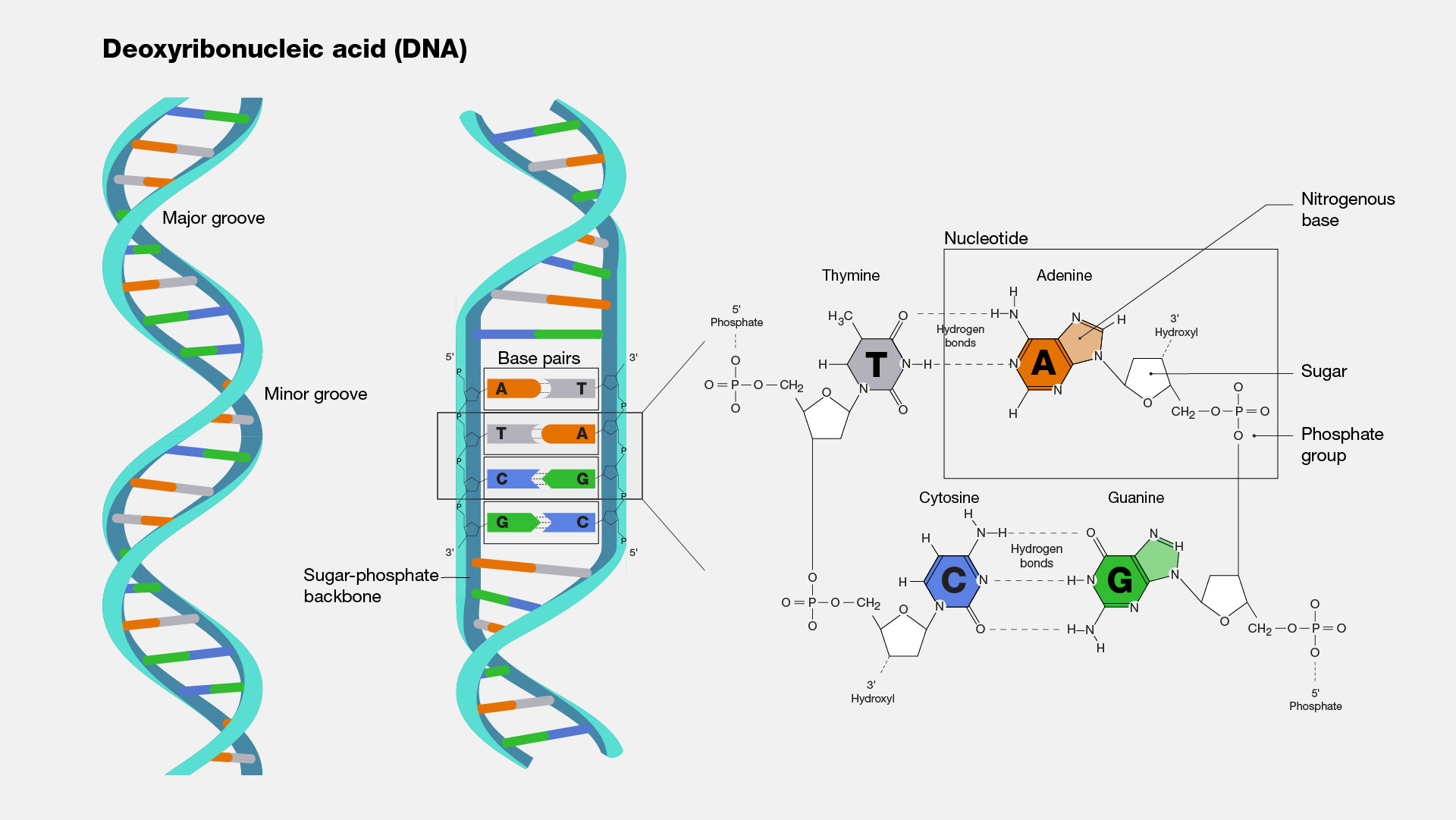
| DNA-(Dexoy-ribo Nucleic Acid) has 4 nucleobases named Adenine, Thymine, Guanine, Cytosine or A, T, G, C. Just like in english we have most basic things: alphabets, in DNA, these nucleobases are most basic things. We need to tokenize them on the basis of these pairs and characters. So this means, our initial vocab is going to be ['A', 'T', 'G', 'C'] instead of 256 utf-8 characters. | |
| Read more about DNA: [Wikipedia/DNA](https://en.wikipedia.org/wiki/DNA) | |
|  | |
| ## Tokenizer: | |
| ### Base Level | |
| It's very basic in working, just like per-character tokenizer which enumerates each and every unique character present in the train file. In our case, we'll have only 4-bases along with '`\n`' and 4-special tokens represented as characters. P, M, U, S as padding, mask, unknown & space token, respectively. | |
| ```python | |
| self.init_vocab = {"\n": 1, "A": 2, "T": 3, "G": 4, "C": 5, "P": 6, "M": 7, "U": 8, "S": 9} | |
| ``` | |
| For encoding and decoding purpose, two functions `string_to_index` & `index_to_string` convert each character into a number from 1 to 9 and decoder takes those 1 to 9 numbers and returns the joint string of respective characters. | |
| ```python | |
| self.string_to_index = {ch: i for i, ch in enumerate(self.chars)} | |
| self.index_to_string = {i: ch for i, ch in enumerate(self.chars)} | |
| ``` | |
| ### K-Mer Tokenization | |
| Let's say we have a long sequence of DNA. This tokenizer splits that sequence into sections of consecutively occurring bases, and each section has length of value equal to `k_mer` which is by default set to 4. | |
| `build_vocab()` function then builds a vocab out of all tokenized sequences by storing them into a new dictionary, seq as key and index as value. And finally, you can save the generated vocab using `save_model()` function and can be loaded later for use. | |
| ```python | |
| tokenizer.load_model('../tokenizer/trained models/base_5k.json') | |
| ``` | |
| I used this tokenizer to train decoder-only model, here is how to use it: | |
| ```python | |
| from tokenizer import KMerTokenizer | |
| tokenizer = KMerTokenizer(k_mers=5) | |
| tokenizer.build_vocab([train_data]) | |
| tokenizer.save_model('../tokenizer/trained models') | |
| encoded_tokens = tokenizer.encode(test_data) | |
| decoded_tokens = tokenizer.decode(encoded_tokens) | |
| ``` | |
| ### Sub-K-Mer Level | |
| It works kind of same as BPE tokenizer, however has some changes in the way it builds its vocab. It first splits it's training into sequences containing only 4 consecutive letters of DNA (same as K-Mer tokenizer with k=4) and then it trains the tokenizer to build new merges based on the frequency of those pairs, like it would have done with the BPE tokenizer. | |
| It can be trained quiet easily and then model file can be saved in two different files; *'.model': contains merges* & *'.json: contains vocab'*. | |
| Encoding and decoding works same as the BPE. | |
| ```python | |
| from tokenizer import KmerPairTokenizer | |
| tokenizer = KmerPairTokenizer() | |
| tokenizer.train(train_data) | |
| tokenizer.save_model('../tokenizer/trained models') | |
| encoded_tokens = tokenizer.encode(test_data) | |
| decoded_tokens = tokenizer.decode(encoded_tokens) | |
| ``` | |
| This tokenizer works fine but it has one problem in decode function, it outputs more tokens than actual present tokens, means: | |
| ```shell | |
| test_data == decoded_tokens is False | |
| ``` | |
| I'll try to fix it and make this work soon, but for now, it's not suitable for use, at-least not for decoding. |